What is an GNN?
GNN Stands for Graph Neural Networks. As its name suggests, GNN is a class of neural networks that operate on graph data, which is a collection of nodes connected by edges. Consider social networks as an example, GNNs can analyze connections to understand communities, predict behavior, or recommend products based on social interactions. Nowadays, GNNs are increasingly used to solve a wide variety of tasks such as model molecules in chemistry, predict protein interactions in biology, or studying citation networks in social science.
Input Data of a GNN:
To start, let`s establish what a graph is. A graph represents the relations (edges) between a collection of entities (nodes). For example, the right panel shows a graph of a chemical compound, from the MUTAG dataset. In this graph, nodes are atoms and edges are bonds between atoms.
Representation
Node Link Diagram or Adjacency Matrix
A graph can be represented as either a node-link diagram or an adjacency matrix. A node-link diagram is an intuitive visual representation of the graph where nodes are depicted as circles and edges are lines connecting the circles. It is easy to understand and informative, especially for small graphs.
An adjacency matrix is a square matrix where the rows and columns represent the nodes, and the value at each intersection (cell) indicates the presence or absence of an edge between the corresponding nodes. It is a compact way to represent the graph, especially for large and dense graphs.
The information of each graph node can be stored as embeddings. For example, in the chemical compound graph, an one-hot encoding is used to represent the atom type, whether it is a carbon, oxygen, nitrogen, etc.
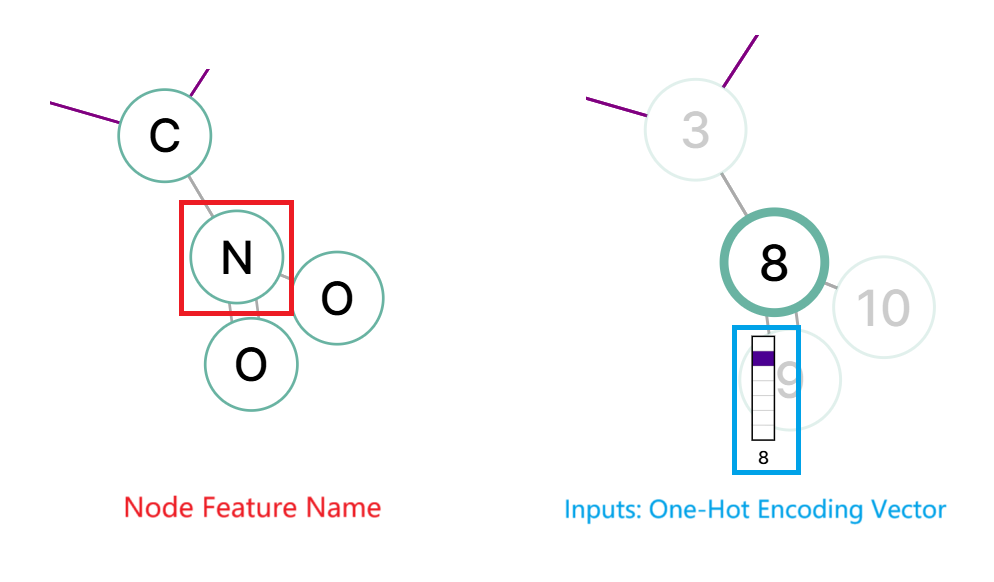 Node Features
Node FeaturesConvolutions on a Graph:
Graphs have an irregular structure can directly use traditional neural networks, which are designed to operate on a fixed, grid-like structure input (such as sentences, images and video). To process graphs, GNNs employ a technique called message passing, where neighboring nodes exchange information and update each other`'`s embeddings to better reflect their interconnectedness and individual features.
Click the "predict" button on the right side and show inner layersMessage-passing forms the backbone of many GNN variants. GNN101 support three popular GNN variants, Graph Convolutional Networks (GCN), Graph Attention Networks (GAT), Graph Sample and Aggregate (GraphSAGE). The main differences between these GNN variants are the way they aggregate information from neighbors and the way they update node embeddings.
We will start with GCN, which is one of the most popular GNN architectures.
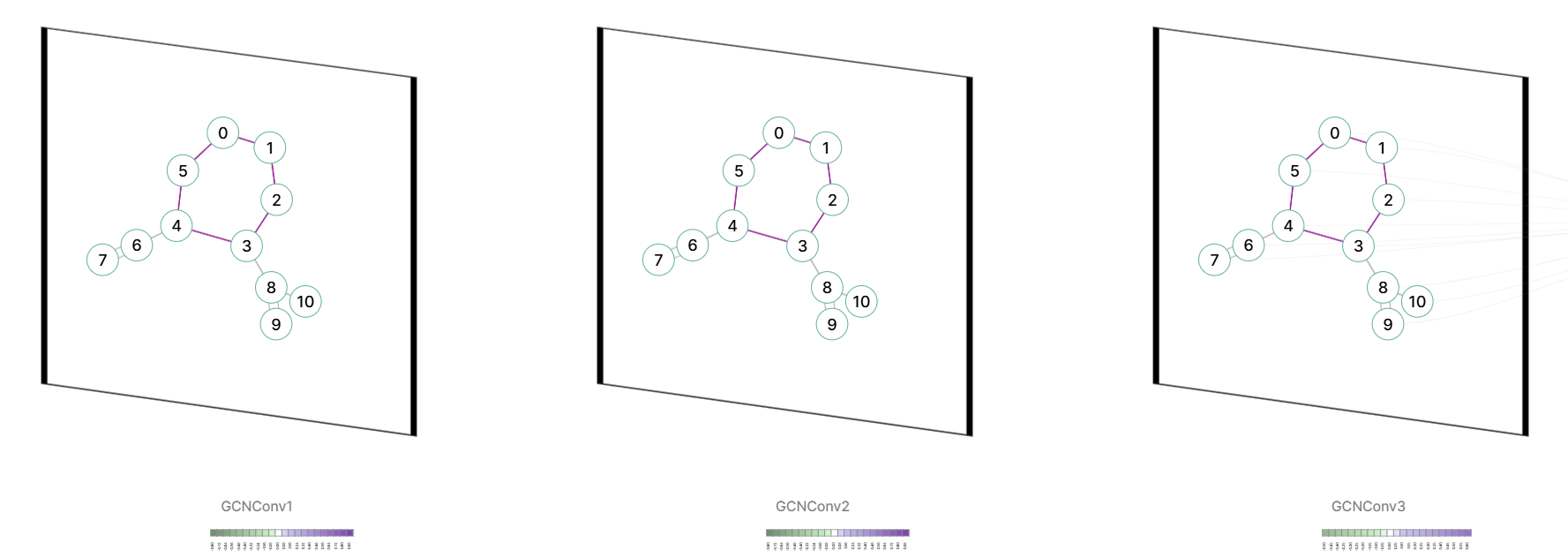 GCNConv
GCNConv- Aggregation with Normalization: First, a node aggregates the feature vectors of its neighbors Xj and itself Xi via a normalized degree matrix W. Intuitively, this reduces the influence of information from nodes with too many neighbors and strengthens the influence from those with fewer neighbors.
- Weighted Transformation: The aggregated information is then transformed by a learnable weight matrix. This matrix allows the model to learn the importance of different features for the central node.
- Activation Function: Finally, we add a bias vector b and a non-linear activation function σ (ReLU) to the aggregated information to obtain an updated feature vector of this node.
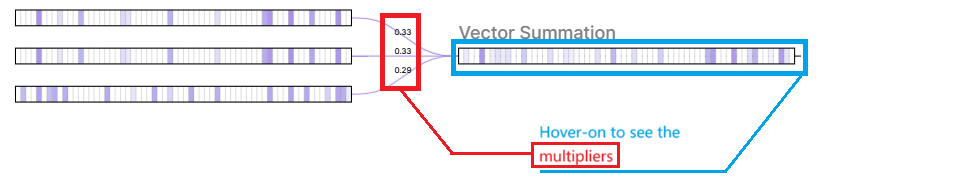 Aggregated Vector
Aggregated Vector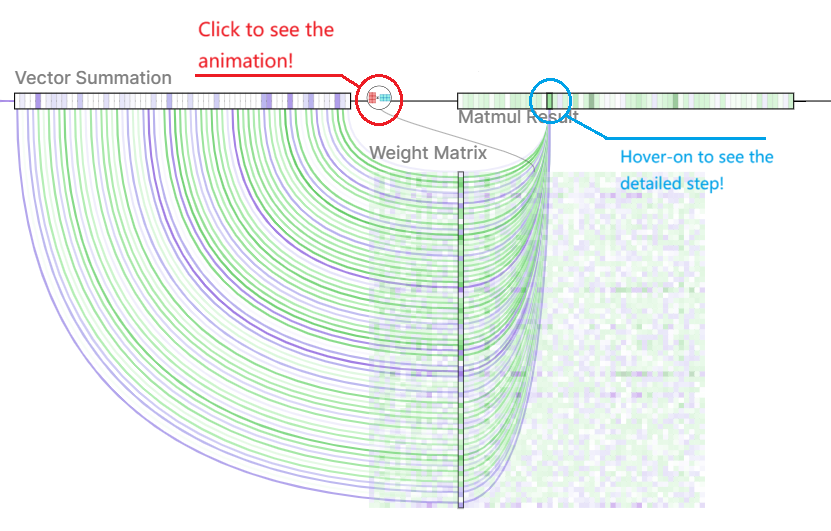 Weighted Transformation
Weighted Transformation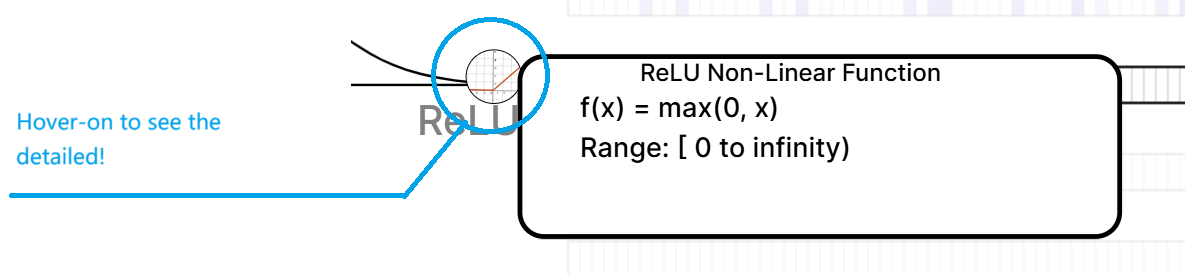 Activation Function
Activation FunctionTasks that GNNs can solve:
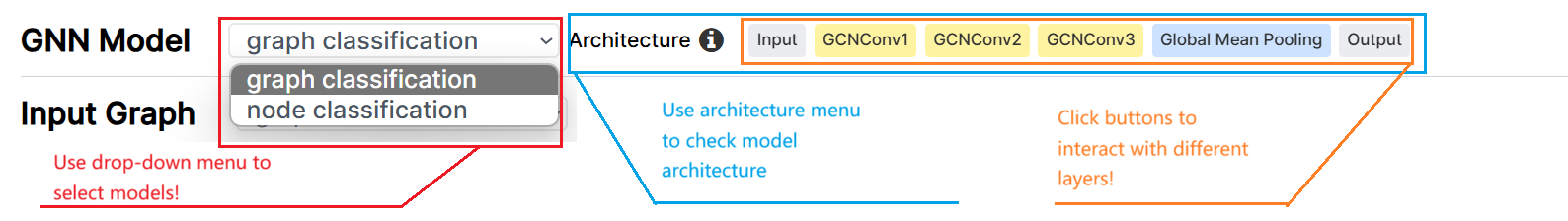 Model Selection Menu
Model Selection MenuBy learning the features of each node, GNNs can then use these node features to solve tasks at different levels of granularity, including graph-level, node-level, community-level, and edge-level tasks. This website currently supports graph-level and node-level tasks. We are working on adding support for other tasks, please stay tuned!
Node-Level TasksGiven the learned features of each node, GNN can directly predict the node properties. For example, in the Karate dataset, the task is to predict the community of each person in the social network. After sevela layers of GCNConv, we apply a fully connected layer to each node to make the prediction.
Graph-Level TasksGNN can predict the properties of the entire graph by aggregating the learned feature of all graph nodes. For example, in the MUTAG dataset, the task is to predict whether a molecule is mutagenic or not. After sevela layers of GCNConv, a global mean pooling layer is used to aggregate the node features into a single graph feature, which is then fed into a fully connected layer to make the prediction.
Edge-Level TasksGNNs can effectively predict edge properties within a graph. In the Twitch dataset, for instance, the model's objective is to determine if two Twitch users are friends. By employing multiple layers, the model combines the features of the two nodes associated with a specific edge using dot product multiplication. Subsequently, a sigmoid function is applied to the resulting score, yielding a probability indicating the likelihood of a friendship between the two users.
About this website:
This website is developed and maintained by the UMN Visual Intelligence Lab. The GNNs you interact with are inferenced real-time on the your web browser, supported by the ONNX web runtime.
🤸 Team Members:- Yilin(Harry) Lu
- Chongwei Chen
- Kexin Huang
- Marinka Zitnik
- Matthew Xu
- Qianwen Wang
💻 Source Code:https://github.com/Visual-Intelligence-UMN/web-gnn-vis
GNN Model
Model-Task
Architecture
Input Graph
Each graph in the MUTAG dataset represents a chemical compound. Nodes are atoms and edges are bonds between atoms. The task is to predict whether a molecule is mutagenic on Salmonella typhimuriumor or not (i.e., can cause genetic mutations in this bacterium or not). The dataset has 188 graphs in total, with 150 graphs in the training set and 38 graphs in the test set.
A graph can be visualized as either a node-link diagram or an adjacency matrix. Hover over the nodes, edges, matrix cells, or matrix labels to highlight their connections.